One thing I always come across when analyzing time-based data is the need to identify events such as changes in signal level, triggers, among others. Many years ago I created a MATLAB function that can find clusters of data values inside a time series. It returns the indexes of the starting point of each cluster, with the corresponding number of samples within each cluster. I have been using the Python version of the function quite a bit recently. Apparently it never gets old!


By doing the proper manipulation of the data, virtually any type of event can be detected and extracted from the time series.
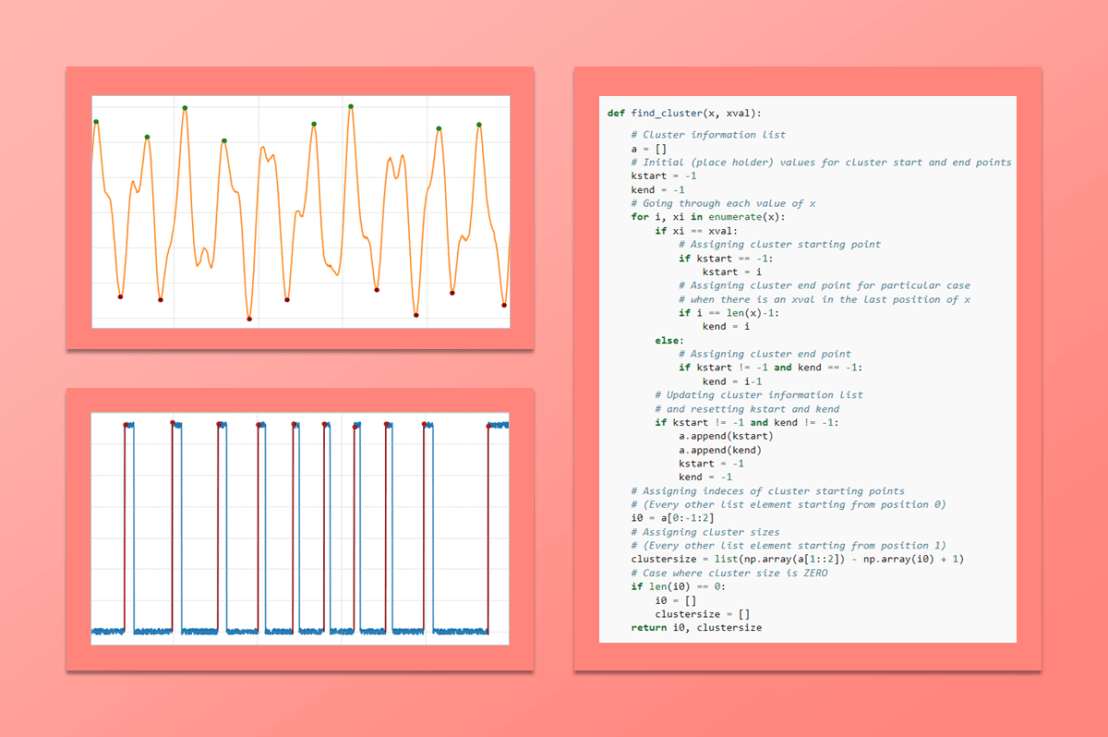
The example on the left shows a generic signal where we search for clusters of data (or events) with values greater than zero. In most cases, it’s good practice to apply a post-processing filter to the signal as the very first step of the data analysis.
The code for the python function used to find the clusters is shown below. Let’s get started by showing some basic usage and then some more interesting cases.
def find_cluster(x, xval):
"""
Find clusters of data in an ndarray that satisfy a certain condition.
:param x: The array containing the data for the cluster search.
:type x: ndarray
:param xval: The value of x that has to be satisfied for clustering.
:type xval: integer, float
:returns: 2-tuple
* i0:
The index of each cluster starting point.
* clustersize:
The corresponding lengths of each cluster.
:rtype: (list, list)
Example
-------
>>> x = np.int32(np.round(np.random.rand(20)+0.1))
>>> i0, clustersize = find_cluster(x, 1)
"""
# Cluster information list
a = []
# Initial (place holder) values for cluster start and end points
kstart = -1
kend = -1
# Going through each value of x
for i, xi in enumerate(x):
if xi == xval:
# Assigning cluster starting point
if kstart == -1:
kstart = i
# Assigning cluster end point for particular case
# when there is an xval in the last position of x
if i == len(x)-1:
kend = i
else:
# Assigning cluster end point
if kstart != -1 and kend == -1:
kend = i-1
# Updating cluster information list
# and resetting kstart and kend
if kstart != -1 and kend != -1:
a.append(kstart)
a.append(kend)
kstart = -1
kend = -1
# Assigning indeces of cluster starting points
# (Every other list element starting from position 0)
i0 = a[0:-1:2]
# Assigning cluster sizes
# (Every other list element starting from position 1)
clustersize = list(np.array(a[1::2]) - np.array(i0) + 1)
# Case where cluster size is ZERO
if len(i0) == 0:
i0 = []
clustersize = []
return i0, clustersize
The example in the doc string above generates a random array x of zeros and ones, in which we use the function to find the clusters of values equal to 1. By running it, one should get something like the results shown below. i0 contains the starting indexes of the clusters of 1s and clustersize contains the number of elements in each cluster.
>>> x = np.int32(np.round(np.random.rand(20)+0.1))
>>> i0, clustersize = find_cluster(x, 1)
>>> x
array([1, 1, 1, 0, 1, 0, 1, 1, 1, 1, 1, 1, 0, 1, 1, 0, 0, 1, 1, 0])
>>> i0
[0, 4, 6, 13, 17]
>>> clustersize
[3, 1, 6, 2, 2]
Finding Signal Peaks

Let’s use the signal from the top of the post and find the signal peaks above 1 and below -1, as illustrated on the left.
The code snippet used to find the peaks as well as the peak values can be seen below. The complete code for it and for all the examples in this post can be found here.
# Finding positive peaks greater than 1
imax = []
icl, ncl = find_cluster(x>1, 1)
for ik, nk in zip(icl, ncl):
imax.append(ik+np.argmax(x[ik:ik+nk])-1)
# Finding negative peaks smaller than 1
imin = []
icl, ncl = find_cluster(x<-1, 1)
for ik, nk in zip(icl, ncl):
imin.append(ik+np.argmin(x[ik:ik+nk])-1)
>>> np.round(x[imax], 2)
array([1.3 , 1.03, 1.47, 1.06, 1.18, 1.53, 1.21, 1.19])
>>> np.round(x[imin], 2)
array([-1.25, -1.19, -1.53, -1.24, -1.07, -1.47, -1.08, -1.3 ])
Finding Trigger Edges

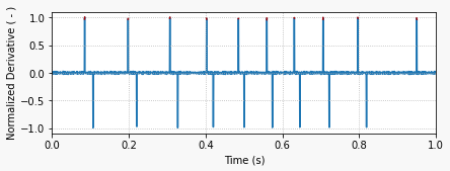
find_cluster can be very useful for determining trigger events in a recorded time series. Unlike most other signals, I usually do not filter a trigger signal, since having a sharp rising edge (at the expense of noise) is critical for a more exact event determination.
The derivative of the trigger signal is then used to determine the rising edges, as shown in the second plot on the left. A threshold value of 0.5 for the derivative (normalized by its maximum value) can be used to locate the trigger edges.
The code snippet below shows how to do it for a fictitious recorded data set.
# Calculating and normalizing trigger signal derivative
dxdt = np.gradient(x, t)
dxdt = dxdt/np.max(dxdt)
# Finding trigger event times using derivative theshold
icl, ncl = find_cluster(dxdt>0.5, 1)
ttrigger = t[icl]
>>> ttrigger
array([0.0845, 0.1975, 0.3065, 0.402 , 0.485 , 0.559 , 0.6305, 0.7055, 0.796 , 0.9495])

One thought on “Event Detection in Signal Processing”